fgeo helps you to install, load, and access the documentation of multiple packages to analyze forest diversity and dynamics (fgeo.analyze, fgeo.plot, fgeo.tool, fgeo.x). This package-collection allows you to manipulate and plot ForestGEO data, and to do common analyses including abundance, demography, and species-habitats associations.
Installation
Make sure your R environment is as follows:
- R version is recent
- All packages are updated (run
update.packages(); maybe useask = FALSE) - No other R session is running
- Current R session is clean (click Session > Restart R)
Install the latest stable version of fgeo from CRAN with:
Or install the development version of fgeo from GitHub with:
Example
library(fgeo)
#> ── Attaching packages ────────────────────────────────────────── fgeo 1.1.3.9000 ──
#> ✔ fgeo.analyze 1.1.10.9000 ✔ fgeo.tool 1.2.5
#> ✔ fgeo.plot 1.1.8 ✔ fgeo.x 1.1.4
#> ── Conflicts ────────────────────────────────────────────────── fgeo_conflicts() ──
#> ✖ fgeo.tool::filter() masks stats::filter()Explore fgeo
On an interactive session, fgeo_help() and fgeo_browse_reference() help you to search documentation.
Access and manipulate data
example_path() allows you to access datasets stored in your R libraries.
example_path()
#> [1] "csv" "mixed_files" "rdata" "rdata_one"
#> [5] "rds" "taxa.csv" "tsv" "vft_4quad.csv"
#> [9] "view" "weird" "xl"
(vft_file <- example_path("view/vft_4quad.csv"))
#> [1] "/home/mauro/R/x86_64-pc-linux-gnu-library/3.6/fgeo.x/extdata/view/vft_4quad.csv"
read_<table>()
read_vft() and read_taxa() import a ViewFullTable and ViewTaxonomy from .tsv or .csv files.
read_vft(vft_file)
#> # A tibble: 500 x 32
#> DBHID PlotName PlotID Family Genus SpeciesName Mnemonic Subspecies
#> <int> <chr> <int> <chr> <chr> <chr> <chr> <chr>
#> 1 385164 luquillo 1 Rubia… Psyc… brachiata PSYBRA <NA>
#> 2 385261 luquillo 1 Urtic… Cecr… schreberia… CECSCH <NA>
#> 3 384600 luquillo 1 Rubia… Psyc… brachiata PSYBRA <NA>
#> 4 608789 luquillo 1 Rubia… Psyc… berteroana PSYBER <NA>
#> 5 388579 luquillo 1 Areca… Pres… acuminata PREMON <NA>
#> 6 384626 luquillo 1 Arali… Sche… morototoni SCHMOR <NA>
#> 7 410958 luquillo 1 Rubia… Psyc… brachiata PSYBRA <NA>
#> 8 385102 luquillo 1 Piper… Piper glabrescens PIPGLA <NA>
#> 9 353163 luquillo 1 Areca… Pres… acuminata PREMON <NA>
#> 10 481018 luquillo 1 Salic… Case… arborea CASARB <NA>
#> # … with 490 more rows, and 24 more variables: SpeciesID <int>,
#> # SubspeciesID <chr>, QuadratName <chr>, QuadratID <int>, PX <dbl>,
#> # PY <dbl>, QX <dbl>, QY <dbl>, TreeID <int>, Tag <chr>, StemID <int>,
#> # StemNumber <int>, StemTag <int>, PrimaryStem <chr>, CensusID <int>,
#> # PlotCensusNumber <int>, DBH <dbl>, HOM <dbl>, ExactDate <date>,
#> # Date <int>, ListOfTSM <chr>, HighHOM <int>, LargeStem <chr>,
#> # Status <chr>
pick_<what>() and drop_<what>()
fgeo is pipe-friendly. You may not use pipes but often they make code easier to read.
Use %>% to emphasize a sequence of actions, rather than the object that the actions are being performed on.
– https://style.tidyverse.org/pipes.html
pick_dbh_under(), drop_status() and friends pick and drop rows from a ForestGEO ViewFullTable or census table.
(census <- fgeo.x::tree5)
#> # A tibble: 30 x 19
#> treeID stemID tag StemTag sp quadrat gx gy MeasureID CensusID
#> <int> <int> <chr> <chr> <chr> <chr> <dbl> <dbl> <int> <int>
#> 1 7624 160987 1089… 175325 TRIP… 722 139. 425. 486675 5
#> 2 8055 10036 1094… 109482 CECS… 522 94.8 424. 468874 5
#> 3 19930 117849 1234… 165576 CASA… 425 61.3 496. 471979 5
#> 4 23746 29677 14473 14473 PREM… 617 100. 328. 442571 5
#> 5 31702 39793 22889 22889 SLOB… 304 53.8 73.8 447307 5
#> 6 35355 44026 27538 27538 SLOB… 1106 203. 110. 449169 5
#> 7 35891 44634 282 282 DACE… 901 172. 14.7 434266 5
#> 8 39705 48888 33371 33370 CASS… 1010 184. 194. 451067 5
#> 9 50184 60798 5830 5830 MATD… 1007 191. 132. 437645 5
#> 10 57380 155867 66962 171649 SLOB… 1414 274. 279. 459427 5
#> # … with 20 more rows, and 9 more variables: dbh <dbl>, pom <chr>,
#> # hom <dbl>, ExactDate <date>, DFstatus <chr>, codes <chr>,
#> # nostems <dbl>, status <chr>, date <dbl>
census %>%
pick_dbh_under(100)
#> # A tibble: 18 x 19
#> treeID stemID tag StemTag sp quadrat gx gy MeasureID CensusID
#> <int> <int> <chr> <chr> <chr> <chr> <dbl> <dbl> <int> <int>
#> 1 7624 160987 1089… 175325 TRIP… 722 139. 425. 486675 5
#> 2 19930 117849 1234… 165576 CASA… 425 61.3 496. 471979 5
#> 3 31702 39793 22889 22889 SLOB… 304 53.8 73.8 447307 5
#> 4 35355 44026 27538 27538 SLOB… 1106 203. 110. 449169 5
#> 5 39705 48888 33371 33370 CASS… 1010 184. 194. 451067 5
#> 6 57380 155867 66962 171649 SLOB… 1414 274. 279. 459427 5
#> 7 95656 129113 1315… 131519 OCOL… 402 79.7 22.8 474157 5
#> 8 96051 129565 1323… 132348 HIRR… 1403 278 40.6 474523 5
#> 9 96963 130553 1347… 134707 TETB… 610 114. 182. 475236 5
#> 10 115310 150789 1652… 165286 MANB… 225 24.0 497. 483175 5
#> 11 121424 158579 1707… 170701 CASS… 811 146. 218. 484785 5
#> 12 121689 158871 1712… 171277 INGL… 515 84.2 285. 485077 5
#> 13 121953 159139 1718… 171809 PSYB… 1318 247. 354. 485345 5
#> 14 124522 162698 1742… 174224 CASS… 1411 279. 210. 488386 5
#> 15 125038 163236 1753… 175335 CASS… 822 153. 426. 488924 5
#> 16 126087 NA 1773… <NA> CASA… 521 89.8 408. NA NA
#> 17 126803 NA 1785… <NA> PSYB… 622 113. 426 NA NA
#> 18 126934 NA 1787… <NA> MICR… 324 47 480. NA NA
#> # … with 9 more variables: dbh <dbl>, pom <chr>, hom <dbl>,
#> # ExactDate <date>, DFstatus <chr>, codes <chr>, nostems <dbl>,
#> # status <chr>, date <dbl>pick_main_stem() and pick_main_stemid() pick the main stem or main stemid(s) of each tree in each census.
add_<column(s)>()
add_status_tree()adds the column status_tree based on the status of all stems of each tree.
stem %>%
select(CensusID, treeID, stemID, status) %>%
add_status_tree()
#> # A tibble: 1,320 x 5
#> CensusID treeID stemID status status_tree
#> <int> <int> <int> <chr> <chr>
#> 1 6 104 143 A A
#> 2 6 119 158 A A
#> 3 NA 180 222 G A
#> 4 NA 180 223 G A
#> 5 6 180 224 G A
#> 6 6 180 225 A A
#> 7 6 602 736 A A
#> 8 6 631 775 A A
#> 9 6 647 793 A A
#> 10 6 1086 1339 A A
#> # … with 1,310 more rowsadd_index() and friends add columns to a ForestGEO-like dataframe.
stem %>%
select(gx, gy) %>%
add_index()
#> Guessing: plotdim = c(320, 500)
#> * If guess is wrong, provide the correct argument `plotdim`
#> # A tibble: 1,320 x 3
#> gx gy index
#> <dbl> <dbl> <dbl>
#> 1 10.3 245. 13
#> 2 183. 410. 246
#> 3 165. 410. 221
#> 4 165. 410. 221
#> 5 165. 410. 221
#> 6 165. 410. 221
#> 7 149. 414. 196
#> 8 38.3 245. 38
#> 9 143. 411. 196
#> 10 68.9 253. 88
#> # … with 1,310 more rowsPlot data
For simplicity, we will focus on only a few species.
autoplot() and friends produce different output depending on the class of input. You can create different input classes, for example, with sp() and sp_elev():
- Use
sp(census)to plot the columnspof acensusdataset – i.e. to plot species distribution.
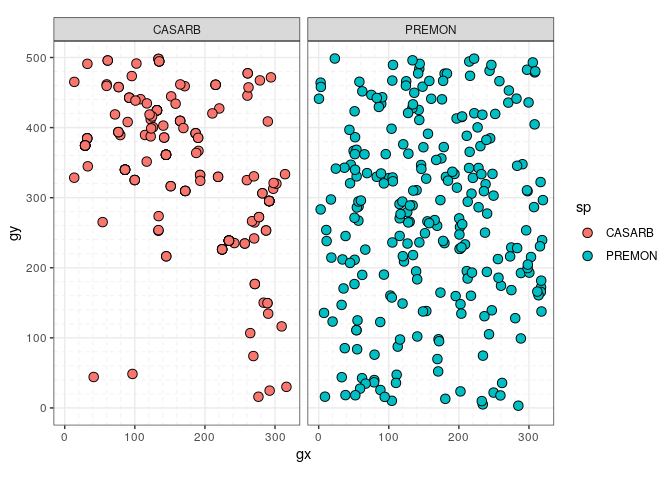
- Use
sp_elev(census, elevation)to plot the columnsspandelevof acensusandelevationdataset, respectively – i.e. to plot species distribution and topography.
elevation <- fgeo.x::elevation
class(sp_elev(stem_2sp, elevation))
#> [1] "sp_elev" "list"
autoplot(sp_elev(stem_2sp, elevation))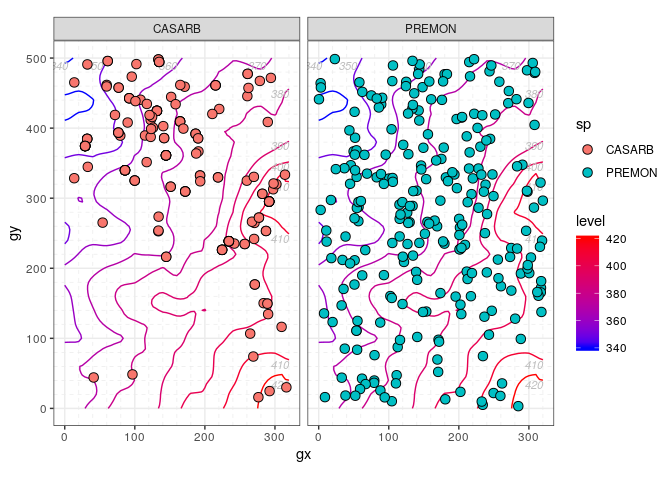
Analyze
Abundance
abundance() and basal_area() calculate abundance and basal area, optionally by groups.
abundance(
pick_main_stem(census)
)
#> # A tibble: 1 x 1
#> n
#> <int>
#> 1 30
by_species <- group_by(census, sp)
basal_area(by_species)
#> # A tibble: 18 x 2
#> # Groups: sp [18]
#> sp basal_area
#> <chr> <dbl>
#> 1 CASARB 437.
#> 2 CASSYL 4146.
#> 3 CECSCH 144150.
#> 4 DACEXC 56832.
#> 5 GUAGUI 9161.
#> 6 HIRRUG 131.
#> 7 INGLAU 141.
#> 8 MANBID 167.
#> 9 MATDOM 45239.
#> 10 MICRAC 0
#> 11 OCOLEU 437.
#> 12 PREMON 78864.
#> 13 PSYBER 0
#> 14 PSYBRA 154.
#> 15 SCHMOR 41187.
#> 16 SLOBER 23377.
#> 17 TETBAL 272.
#> 18 TRIPAL 93.3Demography
recruitment_ctfs(), mortality_ctfs(), and growth_ctfs() calculate recruitment, mortality, and growth. They all output a list. as_tibble() converts the output from a list to a more convenient dataframe.
tree5 <- fgeo.x::tree5
as_tibble(
mortality_ctfs(tree5, tree6)
)
#> Detected dbh ranges:
#> * `census1` = 10.9-323.
#> * `census2` = 10.5-347.
#> Using dbh `mindbh = 0` and above.
#> # A tibble: 1 x 9
#> N D rate lower upper time date1 date2 dbhmean
#> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 27 1 0.00834 0.00195 0.0448 4.52 18938. 20590. 101.Species-habitats association
tt_test() runs a torus translation test to determine habitat associations of tree species. as_tibble() converts the output from a list to a more convenient dataframe. summary() helps you to interpret the result.
# This analysis makes sense only for tree tables
tree <- download_data("luquillo_tree5_random")
habitat <- fgeo.x::habitat
result <- tt_test(tree, habitat)
#> Using `plotdim = c(320, 500)`. To change this value see `?tt_test()`.
#> Using `gridsize = 20`. To change this value see `?tt_test()`.
as_tibble(result)
#> # A tibble: 292 x 8
#> habitat sp N.Hab Gr.Hab Ls.Hab Eq.Hab Rep.Agg.Neut Obs.Quantile
#> * <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 1 ALCFLO 2 1443 153 4 0 0.902
#> 2 2 ALCFLO 1 807 778 15 0 0.504
#> 3 3 ALCFLO 0 0 715 885 -1 0
#> 4 4 ALCFLO 0 0 402 1198 -1 0
#> 5 1 ALCLAT 0 0 544 1056 -1 0
#> 6 2 ALCLAT 1 1432 156 12 0 0.895
#> 7 3 ALCLAT 0 0 324 1276 -1 0
#> 8 4 ALCLAT 0 0 144 1456 -1 0
#> 9 1 ANDINE 1 1117 466 17 0 0.698
#> 10 2 ANDINE 1 1081 510 9 0 0.676
#> # … with 282 more rows
summary(result)
#> # A tibble: 292 x 3
#> sp habitat association
#> <chr> <chr> <chr>
#> 1 ALCFLO 1 neutral
#> 2 ALCFLO 2 neutral
#> 3 ALCFLO 3 repelled
#> 4 ALCFLO 4 repelled
#> 5 ALCLAT 1 repelled
#> 6 ALCLAT 2 neutral
#> 7 ALCLAT 3 repelled
#> 8 ALCLAT 4 repelled
#> 9 ANDINE 1 neutral
#> 10 ANDINE 2 neutral
#> # … with 282 more rowsR code from recent publications by ForestGEO partners
Data have been made available as required by the journal to enable reproduction of the results presented in the paper. Please do not share these data without permission of the ForestGEO plot Principal Investigators (PIs). If you wish to publish papers based on these data, you are also required to get permission from the PIs of the corresponding ForestGEO plots.
- Soil drivers of local-scale tree growth in a lowland tropical forest (Zemunik et al., 2018).
-
Plant diversity increases with the strength of negative density dependence at the global scale (LaManna et al., 2018)
- Response #1: LaManna et al. 2018. Response to Comment on “Plant diversity increases with the strength of negative density dependence at the global scale” Science Vol. 360, Issue 6391, eaar3824. DOI: 10.1126/science.aar3824
- Response #2: LaManna et al. 2018. Response to Comment on “Plant diversity increases with the strength of negative density dependence at the global scale”. Science Vol. 360, Issue 6391, eaar5245. DOI: 10.1126/science.aar5245
Acknowledgments
Thanks to all partners of ForestGEO for sharing their ideas and code. For feedback on fgeo, special thanks to Gabriel Arellano, Stuart Davies, Lauren Krizel, Sean McMahon, and Haley Overstreet. For all other help, I thank contributors in the the documentation of the features they helped with.
 Analyze forest diversity and dynamics
Analyze forest diversity and dynamics