Krige soil data
Based on code developed by Graham Zemunik
Mauro Lepore
2020-03-11
Source:vignettes/siteonly/krige.Rmd
krige.RmdThis article shows how to krig soil data with the function krig() developed by Graham Zemunik (see ?krig()).
library(ggplot2)
library(purrr)
library(fgeo.tool)
#>
#> Attaching package: 'fgeo.tool'
#> The following object is masked from 'package:stats':
#>
#> filter
library(fgeo.krig)
#> Warning in fun(libname, pkgname): couldn't connect to display ":99.0"soil <- fgeo.krig::soil_random
str(soil)
#> Classes 'tbl_df', 'tbl' and 'data.frame': 100 obs. of 3 variables:
#> $ gx : int 9 9 29 29 29 49 69 69 69 89 ...
#> $ gy : int 110 270 130 290 370 390 90 130 330 190 ...
#> $ m3al: num 927 716 809 1115 419 ...Compare parameters
Krige with automated parameters.
auto <- krig(soil, var = "m3al", quiet = TRUE)
#> Guessing: plotdim = c(1000, 500)
summary(auto)
#> var: m3al
#> df
#> Classes 'tbl_df', 'tbl' and 'data.frame': 1250 obs. of 3 variables:
#> $ x: num 10 30 50 70 90 110 130 150 170 190 ...
#> $ y: num 10 10 10 10 10 10 10 10 10 10 ...
#> $ z: num 950 944 939 933 928 ...
#>
#> df.poly
#> Classes 'tbl_df', 'tbl' and 'data.frame': 1250 obs. of 3 variables:
#> $ gx: num 10 30 50 70 90 110 130 150 170 190 ...
#> $ gy: num 10 10 10 10 10 10 10 10 10 10 ...
#> $ z : num 949 943 938 933 927 ...
#>
#> lambda
#> 'numeric'
#> num 1
#>
#> vg
#> 'variogram'
#> List of 20
#> $ u : num [1:12] 30.3 42.9 51.1 60.9 86.5 ...
#> $ v : num [1:12] 60522 54239 174166 55226 44193 ...
#> $ n : num [1:12] 21 25 5 81 94 57 155 167 247 288 ...
#> $ sd : num [1:12] 96683 58618 214627 60326 52526 ...
#> $ bins.lim : num [1:31] 1.00e-12 2.00 2.38 2.84 3.38 ...
#> $ ind.bin : logi [1:30] FALSE FALSE FALSE FALSE FALSE FALSE ...
#> $ var.mark : num 55039
#> $ beta.ols : num 2.62e-08
#> $ output.type : chr "bin"
#> $ max.dist : num 320
#> $ estimator.type : chr "classical"
#> $ n.data : int 100
#> $ lambda : num 1
#> $ trend : chr "cte"
#> $ pairs.min : num 5
#> $ nugget.tolerance: num 1e-12
#> $ direction : chr "omnidirectional"
#> $ tolerance : chr "none"
#> $ uvec : num [1:30] 1 2.19 2.61 3.11 3.7 ...
#> $ call : language variog(geodata = geodata, breaks = breaks, trend = trend, pairs.min = 5)
#>
#> vm
#> 'variomodel', variofit'
#> List of 17
#> $ nugget : num 56634
#> $ cov.pars : num [1:2] 87783 2105
#> $ cov.model : chr "gaussian"
#> $ kappa : num 0.5
#> $ value : num 1.49e+11
#> $ trend : chr "cte"
#> $ beta.ols : num 2.62e-08
#> $ practicalRange : num 3644
#> $ max.dist : num 320
#> $ minimisation.function: chr "optim"
#> $ weights : chr "npairs"
#> $ method : chr "WLS"
#> $ fix.nugget : logi FALSE
#> $ fix.kappa : logi TRUE
#> $ lambda : num 1
#> $ message : chr "optim convergence code: 0"
#> $ call : language variofit(vario = vg, ini.cov.pars = c(initialVal, startRange), cov.model = varModels[i], nugget = initialVal)as_tibble(auto)
#> # A tibble: 1,250 x 4
#> var x y z
#> <chr> <dbl> <dbl> <dbl>
#> 1 m3al 10 10 950.
#> 2 m3al 30 10 944.
#> 3 m3al 50 10 939.
#> 4 m3al 70 10 933.
#> 5 m3al 90 10 928.
#> 6 m3al 110 10 922.
#> 7 m3al 130 10 917.
#> 8 m3al 150 10 911.
#> 9 m3al 170 10 906.
#> 10 m3al 190 10 900.
#> # … with 1,240 more rowsTry also:

With custom parameters (arbitrary but based on automated kriging parameters).
params <- list(
model = "circular", range = 100, nugget = 1000, sill = 46000, kappa = 0.5
)
custom <- krig(soil, var = "m3al", params = params, quiet = TRUE)
#> Guessing: plotdim = c(1000, 500)Compare.
title_auto <- "Automated parameters"
ggplot(as_tibble(auto), aes(x = x, y = y, fill = z)) +
geom_tile() +
coord_equal() +
labs(title = title_auto)
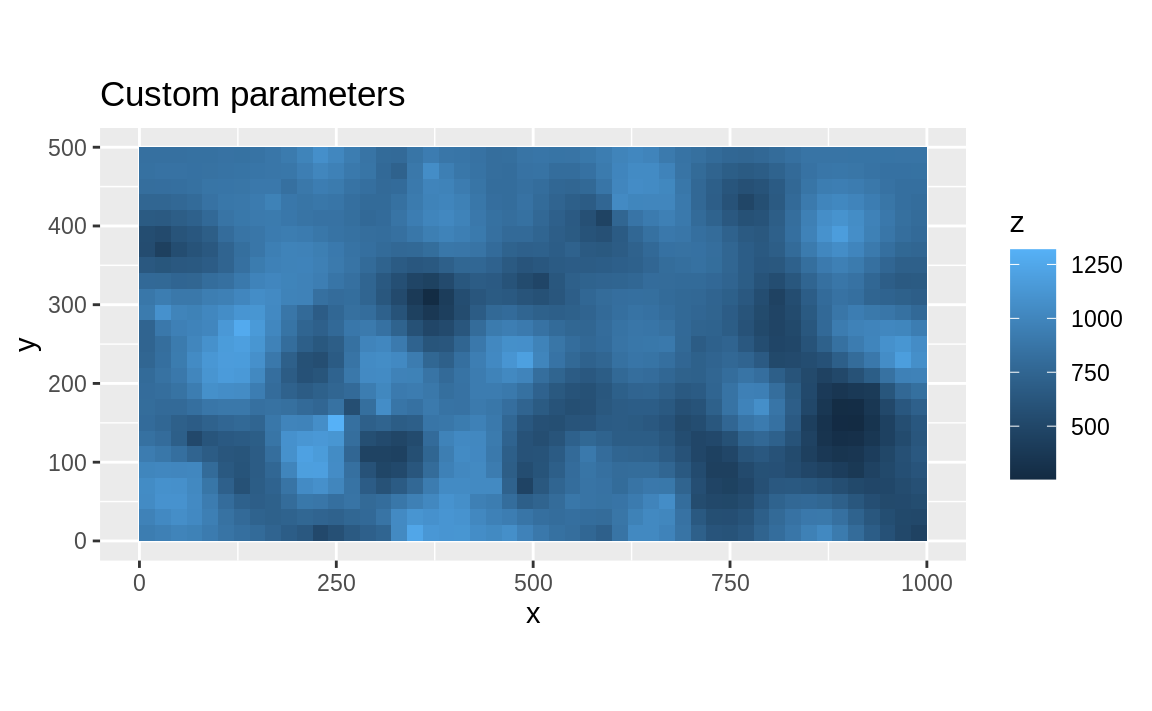
title_custom <- "Custom parameters"
ggplot(as_tibble(custom), aes(x = x, y = y, fill = z)) +
geom_tile() +
coord_equal() +
labs(title = title_custom)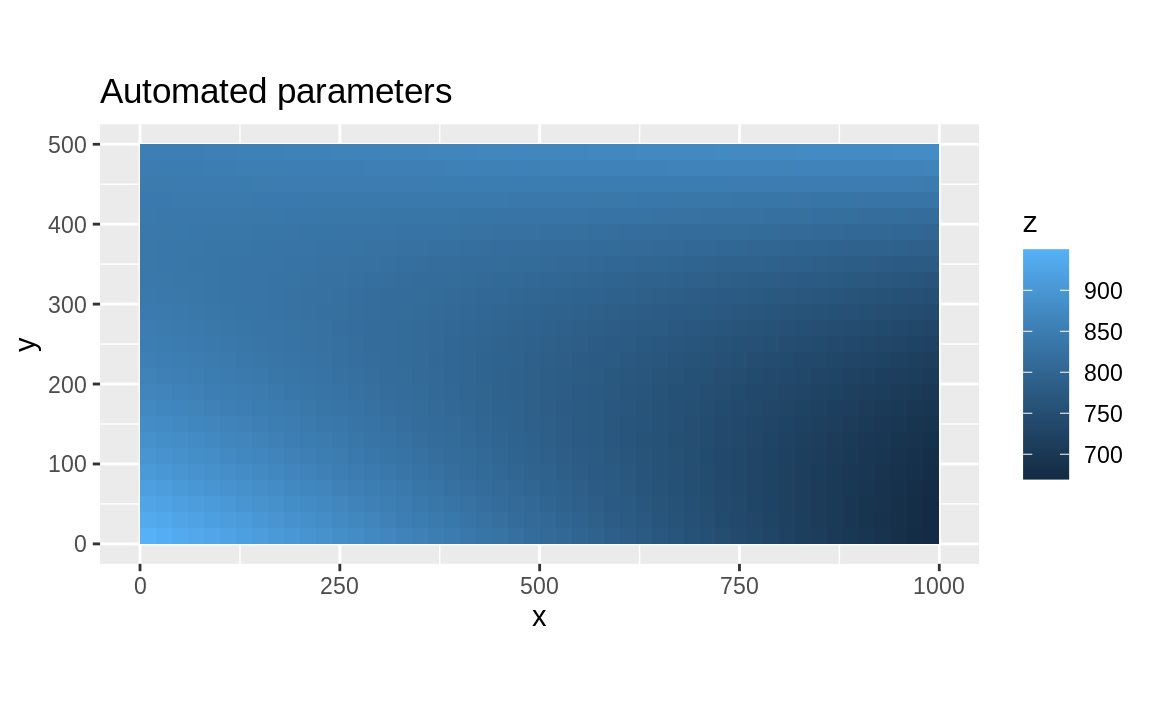
Iterate over multiple soil variables
soil <- fgeo.krig::soil_fake
soil
#> # A tibble: 30 x 5
#> gx gy mg c p
#> <int> <int> <dbl> <dbl> <dbl>
#> 1 40 193 0.67 1.75 6.5
#> 2 56 30 0.5 2.25 5.90
#> 3 61 102 0.65 2.05 6.40
#> 4 67 110 0.5 2.35 6.40
#> 5 113 16 0.56 1.45 6.20
#> 6 173 134 0.74 1.55 7
#> 7 239 252 0.47 0.85 5.90
#> 8 257 442 0.52 2.25 6.20
#> 9 283 288 0.45 0.45 6.7
#> 10 294 181 0.7 2.45 7.1
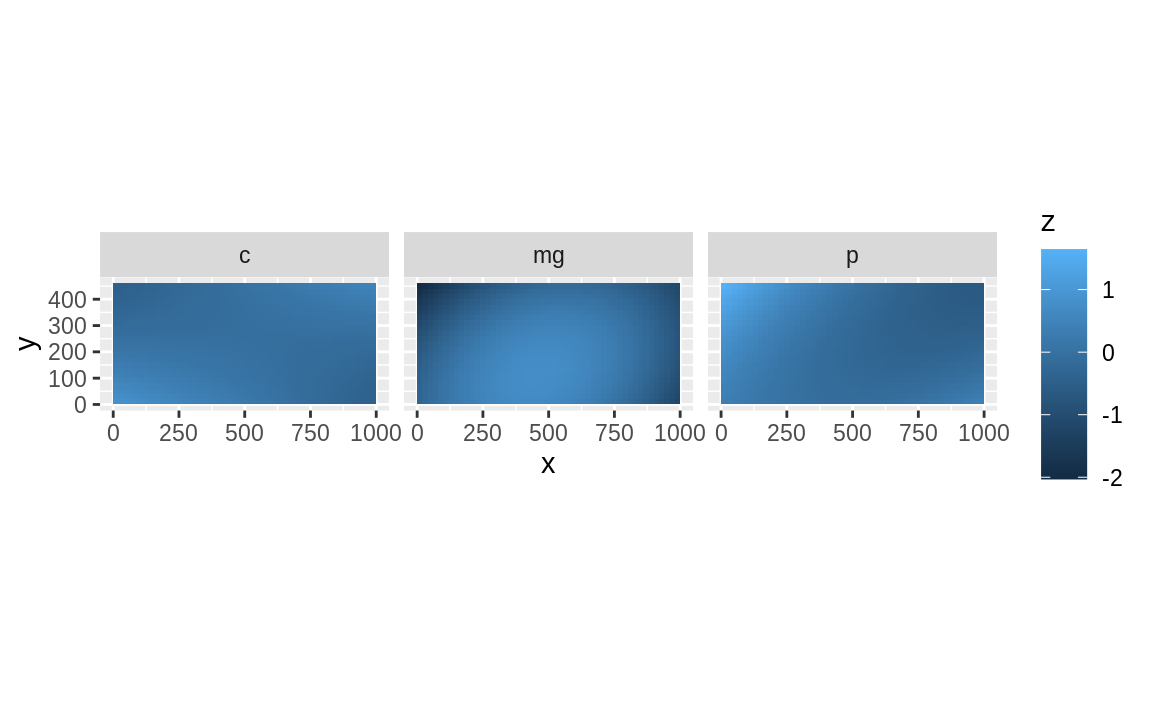
#> # … with 20 more rowsScale soil data to make their values fall within the same range. This is not crucial, but it will ease visual comparison.
soil_vars <- c("mg", "c", "p")
soil_scaled <- modify_at(soil_fake, .at = soil_vars, scale)
krig_list <- krig(soil_scaled, soil_vars, quiet = TRUE)
#> Guessing: plotdim = c(1000, 460)
summary(krig_list)
#> var: mg
#> df
#> Classes 'tbl_df', 'tbl' and 'data.frame': 1150 obs. of 3 variables:
#> $ x: num 10 30 50 70 90 110 130 150 170 190 ...
#> $ y: num 10 10 10 10 10 10 10 10 10 10 ...
#> $ z: num -0.3186 -0.2256 -0.1372 -0.0535 0.0256 ...
#>
#> df.poly
#> Classes 'tbl_df', 'tbl' and 'data.frame': 1150 obs. of 3 variables:
#> $ gx: num 10 30 50 70 90 110 130 150 170 190 ...
#> $ gy: num 10 10 10 10 10 10 10 10 10 10 ...
#> $ z : num -0.3186 -0.2256 -0.1372 -0.0535 0.0256 ...
#>
#> lambda
#> 'numeric'
#> num 1
#>
#> vg
#> 'variogram'
#> List of 20
#> $ u : num [1:9] 60.9 86.5 103 122.7 146.1 ...
#> $ v : num [1:9] 0.858 0.852 1.19 0.856 1.229 ...
#> $ n : num [1:9] 7 9 10 10 18 19 36 34 38
#> $ sd : num [1:9] 1.204 0.981 1.444 0.835 0.996 ...
#> $ bins.lim : num [1:31] 1.00e-12 2.00 2.38 2.84 3.38 ...
#> $ ind.bin : logi [1:30] FALSE FALSE FALSE FALSE FALSE FALSE ...
#> $ var.mark : num 0.819
#> $ beta.ols : num -5.08e-09
#> $ output.type : chr "bin"
#> $ max.dist : num 320
#> $ estimator.type : chr "classical"
#> $ n.data : int 30
#> $ lambda : num 1
#> $ trend : chr "cte"
#> $ pairs.min : num 5
#> $ nugget.tolerance: num 1e-12
#> $ direction : chr "omnidirectional"
#> $ tolerance : chr "none"
#> $ uvec : num [1:30] 1 2.19 2.61 3.11 3.7 ...
#> $ call : language variog(geodata = geodata, breaks = breaks, trend = trend, pairs.min = 5)
#>
#> vm
#> 'variomodel', variofit'
#> List of 17
#> $ nugget : num 0.931
#> $ cov.pars : num [1:2] 0 160
#> $ cov.model : chr "gaussian"
#> $ kappa : num 0.5
#> $ value : num 19.9
#> $ trend : chr "cte"
#> $ beta.ols : num -5.08e-09
#> $ practicalRange : num 277
#> $ max.dist : num 320
#> $ minimisation.function: chr "optim"
#> $ weights : chr "npairs"
#> $ method : chr "WLS"
#> $ fix.nugget : logi FALSE
#> $ fix.kappa : logi TRUE
#> $ lambda : num 1
#> $ message : chr "optim convergence code: 0"
#> $ call : language variofit(vario = vg, ini.cov.pars = c(initialVal, startRange), cov.model = varModels[i], nugget = initialVal)
#>
#> var: c
#> df
#> Classes 'tbl_df', 'tbl' and 'data.frame': 1150 obs. of 3 variables:
#> $ x: num 10 30 50 70 90 110 130 150 170 190 ...
#> $ y: num 10 10 10 10 10 10 10 10 10 10 ...
#> $ z: num 0.837 0.811 0.785 0.759 0.733 ...
#>
#> df.poly
#> Classes 'tbl_df', 'tbl' and 'data.frame': 1150 obs. of 3 variables:
#> $ gx: num 10 30 50 70 90 110 130 150 170 190 ...
#> $ gy: num 10 10 10 10 10 10 10 10 10 10 ...
#> $ z : num 0.837 0.811 0.785 0.759 0.733 ...
#>
#> lambda
#> 'numeric'
#> num 1
#>
#> vg
#> 'variogram'
#> List of 20
#> $ u : num [1:9] 60.9 86.5 103 122.7 146.1 ...
#> $ v : num [1:9] 0.834 1.239 2.593 1.597 0.621 ...
#> $ n : num [1:9] 7 9 10 10 18 19 36 34 38
#> $ sd : num [1:9] 1.22 1.41 1.86 1.47 1.19 ...
#> $ bins.lim : num [1:31] 1.00e-12 2.00 2.38 2.84 3.38 ...
#> $ ind.bin : logi [1:30] FALSE FALSE FALSE FALSE FALSE FALSE ...
#> $ var.mark : num 0.931
#> $ beta.ols : num 1.77e-11
#> $ output.type : chr "bin"
#> $ max.dist : num 320
#> $ estimator.type : chr "classical"
#> $ n.data : int 30
#> $ lambda : num 1
#> $ trend : chr "cte"
#> $ pairs.min : num 5
#> $ nugget.tolerance: num 1e-12
#> $ direction : chr "omnidirectional"
#> $ tolerance : chr "none"
#> $ uvec : num [1:30] 1 2.19 2.61 3.11 3.7 ...
#> $ call : language variog(geodata = geodata, breaks = breaks, trend = trend, pairs.min = 5)
#>
#> vm
#> 'variomodel', variofit'
#> List of 17
#> $ nugget : num 1.04
#> $ cov.pars : num [1:2] 0 161
#> $ cov.model : chr "exponential"
#> $ kappa : num 0.5
#> $ value : num 40.1
#> $ trend : chr "cte"
#> $ beta.ols : num 1.77e-11
#> $ practicalRange : num 481
#> $ max.dist : num 320
#> $ minimisation.function: chr "optim"
#> $ weights : chr "npairs"
#> $ method : chr "WLS"
#> $ fix.nugget : logi FALSE
#> $ fix.kappa : logi TRUE
#> $ lambda : num 1
#> $ message : chr "optim convergence code: 0"
#> $ call : language variofit(vario = vg, ini.cov.pars = c(initialVal, startRange), cov.model = varModels[i], nugget = initialVal)
#>
#> var: p
#> df
#> Classes 'tbl_df', 'tbl' and 'data.frame': 1150 obs. of 3 variables:
#> $ x: num 10 30 50 70 90 110 130 150 170 190 ...
#> $ y: num 10 10 10 10 10 10 10 10 10 10 ...
#> $ z: num 0.375 0.337 0.301 0.266 0.233 ...
#>
#> df.poly
#> Classes 'tbl_df', 'tbl' and 'data.frame': 1150 obs. of 3 variables:
#> $ gx: num 10 30 50 70 90 110 130 150 170 190 ...
#> $ gy: num 10 10 10 10 10 10 10 10 10 10 ...
#> $ z : num 0.375 0.337 0.301 0.266 0.233 ...
#>
#> lambda
#> 'numeric'
#> num 1
#>
#> vg
#> 'variogram'
#> List of 20
#> $ u : num [1:9] 60.9 86.5 103 122.7 146.1 ...
#> $ v : num [1:9] 1.311 1.325 0.366 1.332 1.275 ...
#> $ n : num [1:9] 7 9 10 10 18 19 36 34 38
#> $ sd : num [1:9] 1.963 1.371 0.441 1.355 1.616 ...
#> $ bins.lim : num [1:31] 1.00e-12 2.00 2.38 2.84 3.38 ...
#> $ ind.bin : logi [1:30] FALSE FALSE FALSE FALSE FALSE FALSE ...
#> $ var.mark : num 0.886
#> $ beta.ols : num -1.19e-09
#> $ output.type : chr "bin"
#> $ max.dist : num 320
#> $ estimator.type : chr "classical"
#> $ n.data : int 30
#> $ lambda : num 1
#> $ trend : chr "cte"
#> $ pairs.min : num 5
#> $ nugget.tolerance: num 1e-12
#> $ direction : chr "omnidirectional"
#> $ tolerance : chr "none"
#> $ uvec : num [1:30] 1 2.19 2.61 3.11 3.7 ...
#> $ call : language variog(geodata = geodata, breaks = breaks, trend = trend, pairs.min = 5)
#>
#> vm
#> 'variomodel', variofit'
#> List of 17
#> $ nugget : num 1.01
#> $ cov.pars : num [1:2] 0 160
#> $ cov.model : chr "circular"
#> $ kappa : num 0.5
#> $ value : num 8.99
#> $ trend : chr "cte"
#> $ beta.ols : num -1.19e-09
#> $ practicalRange : num 160
#> $ max.dist : num 320
#> $ minimisation.function: chr "optim"
#> $ weights : chr "npairs"
#> $ method : chr "WLS"
#> $ fix.nugget : logi FALSE
#> $ fix.kappa : logi TRUE
#> $ lambda : num 1
#> $ message : chr "optim convergence code: 0"
#> $ call : language variofit(vario = vg, ini.cov.pars = c(initialVal, startRange), cov.model = varModels[i], nugget = initialVal)Transform krig results to a tibble for easier manipulation and visualization.
# as.data.frame(krig_list) also works but is hader to visualize
(krig_df <- as_tibble(krig_list))
#> # A tibble: 3,450 x 4
#> var x y z
#> <chr> <dbl> <dbl> <dbl>
#> 1 mg 10 10 -0.319
#> 2 mg 30 10 -0.226
#> 3 mg 50 10 -0.137
#> 4 mg 70 10 -0.0535
#> 5 mg 90 10 0.0256
#> 6 mg 110 10 0.100
#> 7 mg 130 10 0.170
#> 8 mg 150 10 0.235
#> 9 mg 170 10 0.296
#> 10 mg 190 10 0.352
#> # … with 3,440 more rows
tail(krig_df)
#> # A tibble: 6 x 4
#> var x y z
#> <chr> <dbl> <dbl> <dbl>
#> 1 p 890 450 -0.598
#> 2 p 910 450 -0.613
#> 3 p 930 450 -0.626
#> 4 p 950 450 -0.639
#> 5 p 970 450 -0.649
#> 6 p 990 450 -0.658